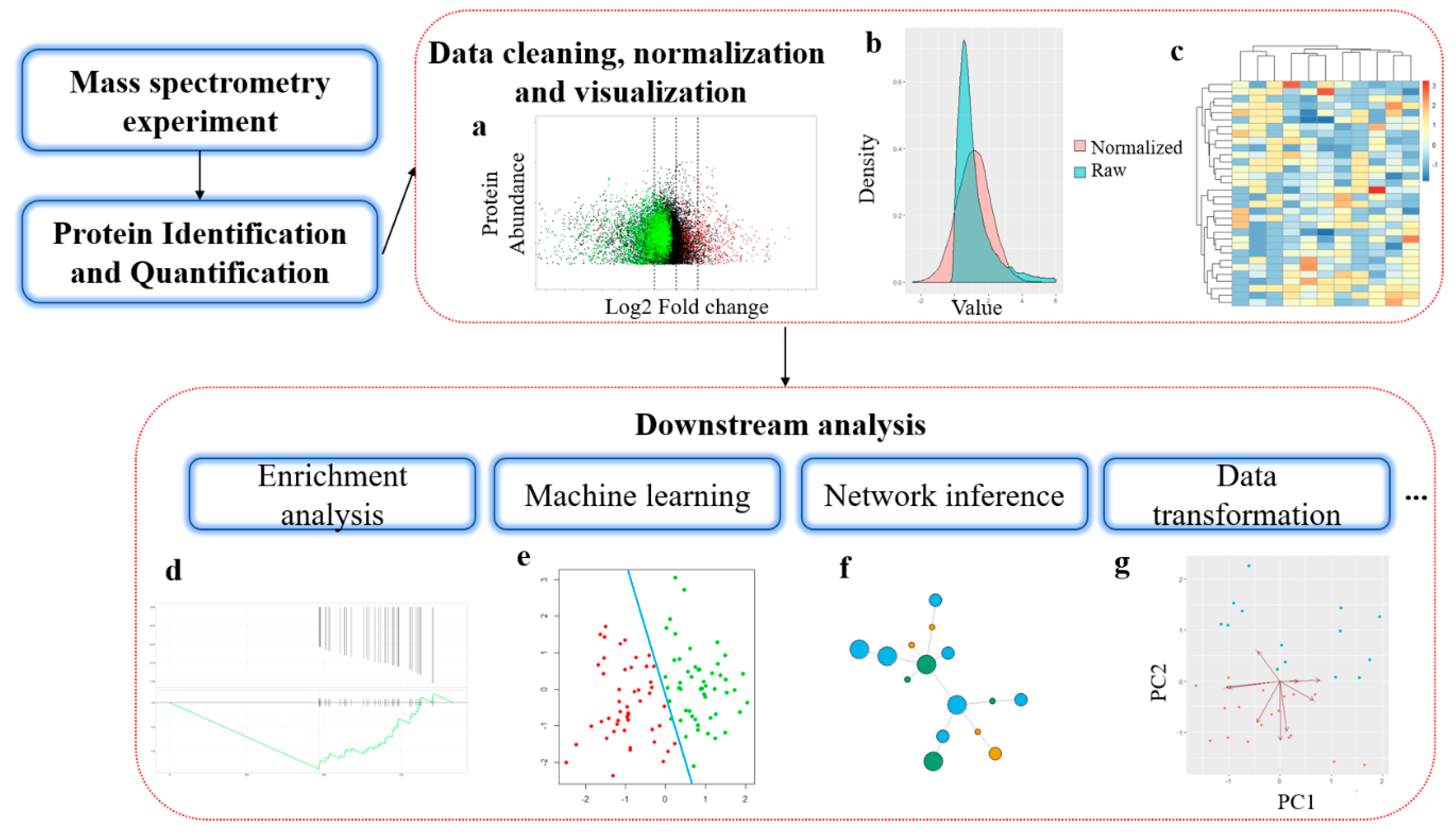
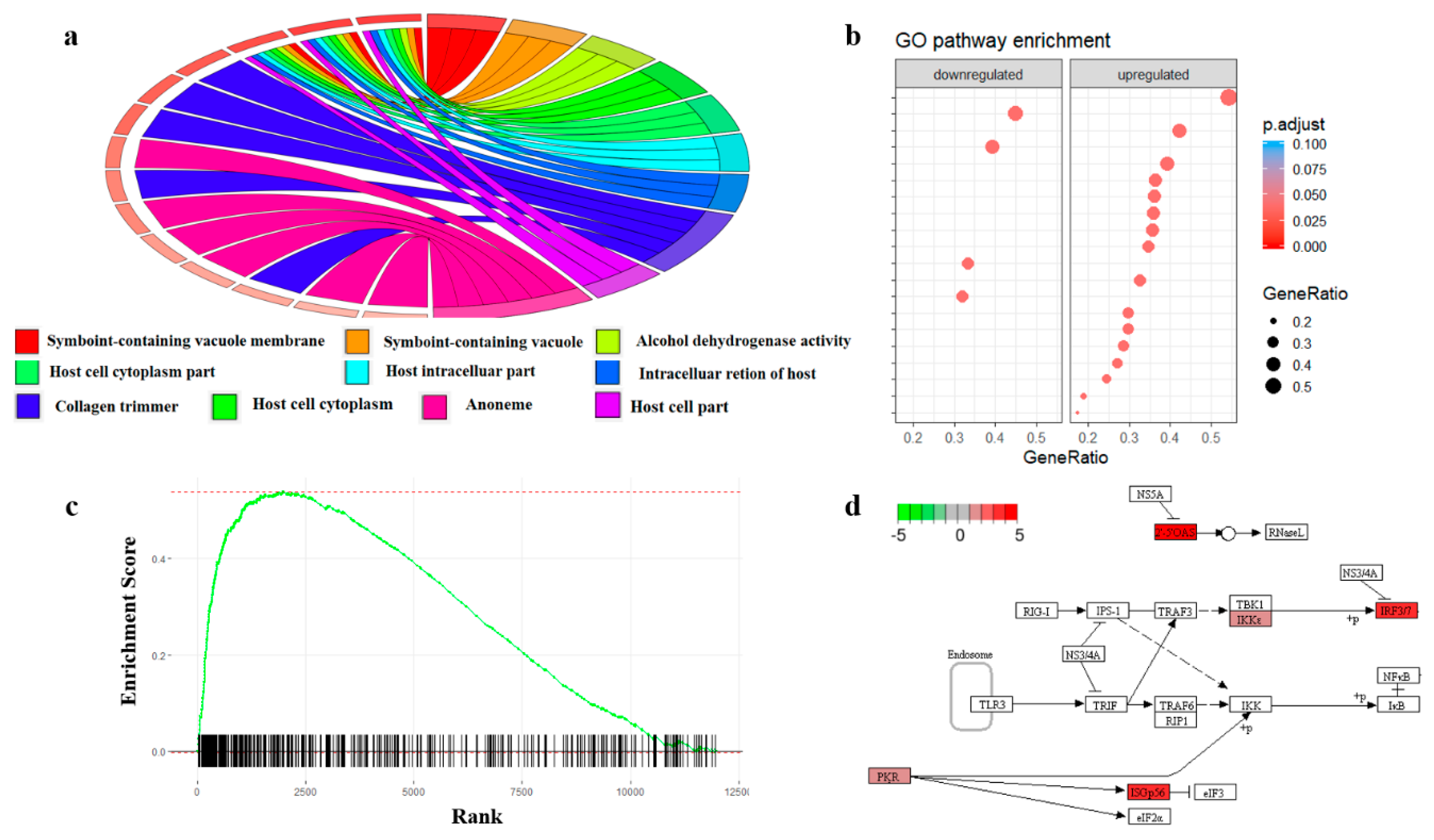
Label-free proteomics data clustering of heart tissue from control (C)... | Download Scientific Diagram

Visualization of proteomics data using R and Bioconductor - Gatto - 2015 - PROTEOMICS - Wiley Online Library

IonStar enables high-precision, low-missing-data proteomics quantification in large biological cohorts | PNAS
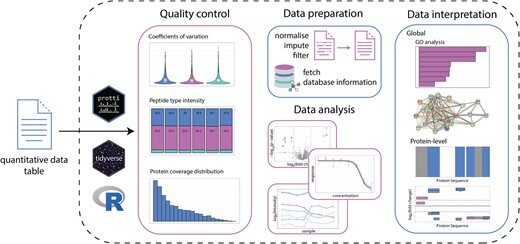
An R package for comprehensive data analysis of peptide- and protein-centric bottom-up proteomics data

Genoppi is an open-source software for robust and standardized integration of proteomic and genetic data | Nature Communications
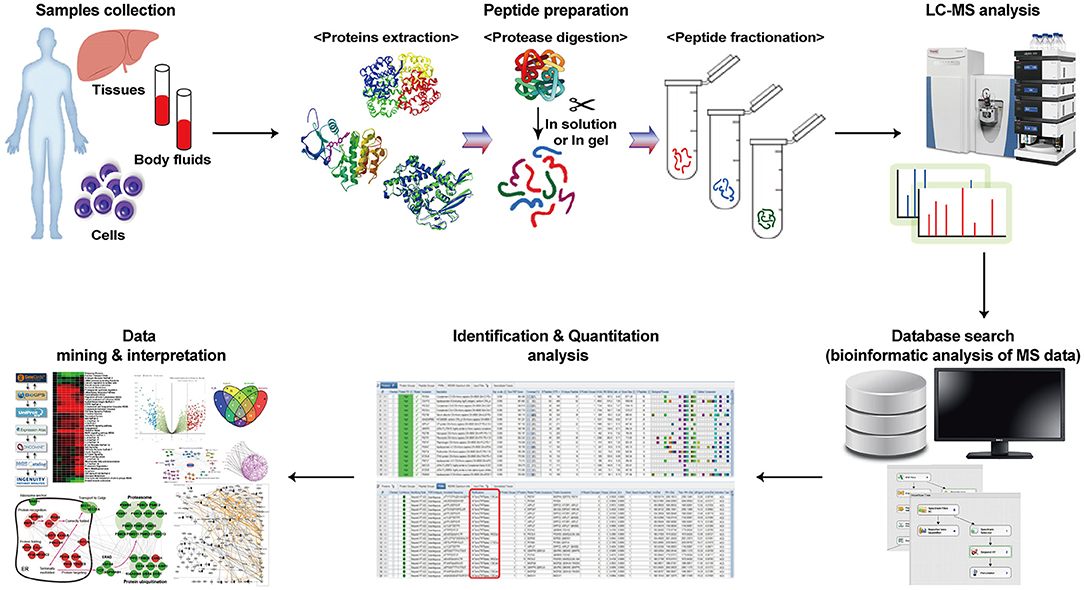
Frontiers | Application of Proteomics in Cancer: Recent Trends and Approaches for Biomarkers Discovery

Comparison of Database Searching Programs for the Analysis of Single-Cell Proteomics Data | Journal of Proteome Research
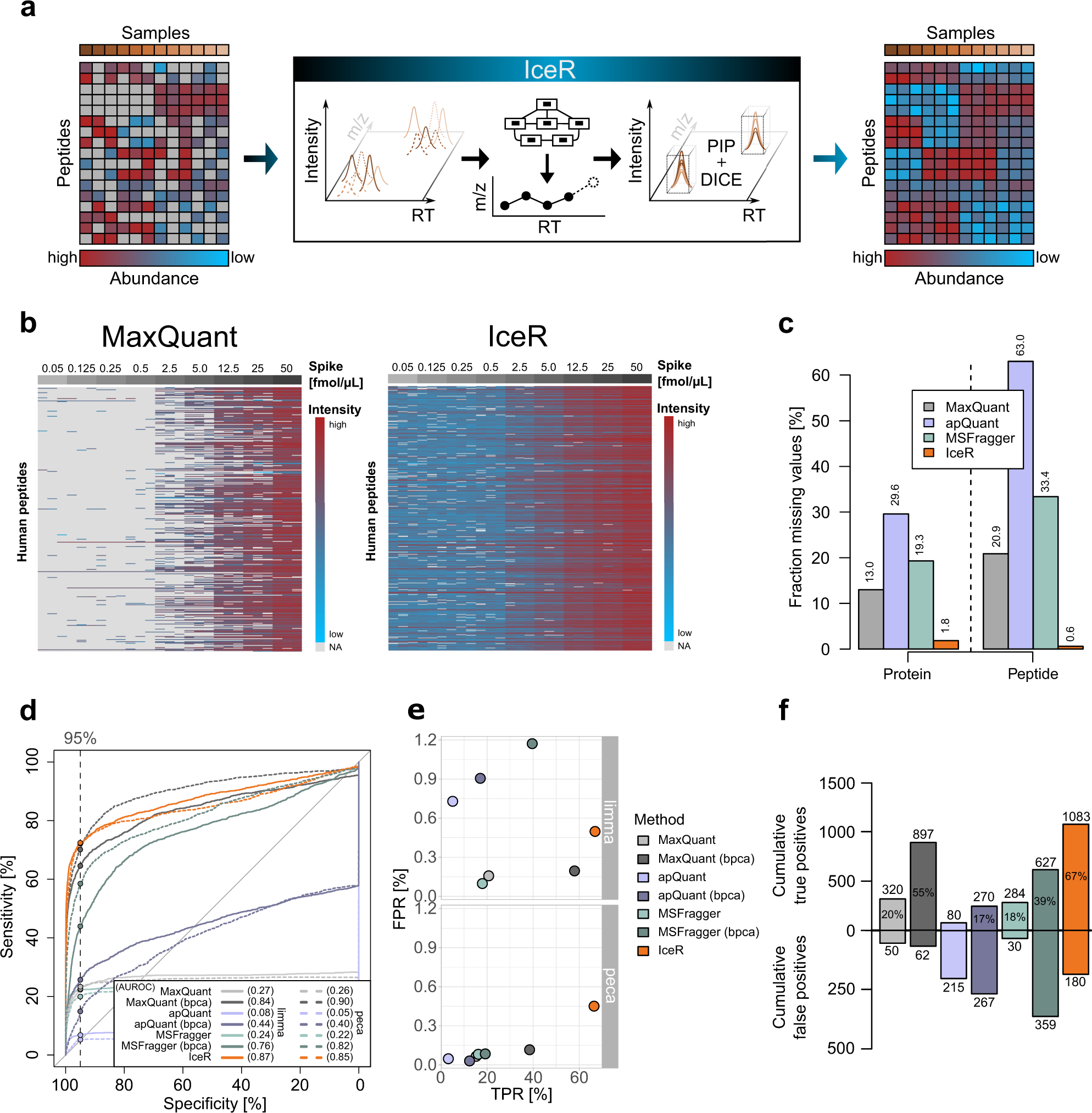
IceR improves proteome coverage and data completeness in global and single-cell proteomics | Nature Communications
![PDF] ProVision: a web-based platform for rapid analysis of proteomics data processed by MaxQuant | Semantic Scholar PDF] ProVision: a web-based platform for rapid analysis of proteomics data processed by MaxQuant | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/2e41862f6ae4d12ef81d7f777fc8819ac8bd6146/2-Figure1-1.png)
PDF] ProVision: a web-based platform for rapid analysis of proteomics data processed by MaxQuant | Semantic Scholar
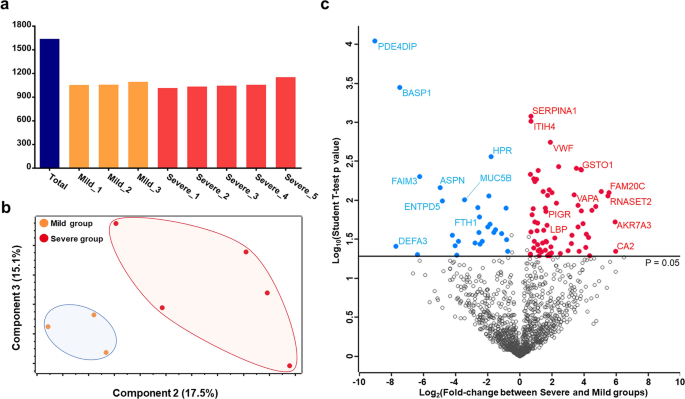
In-depth blood proteome profiling analysis revealed distinct functional characteristics of plasma proteins between severe and non-severe COVID-19 patients | Scientific Reports

DIAproteomics: A Multifunctional Data Analysis Pipeline for Data-Independent Acquisition Proteomics and Peptidomics | Journal of Proteome Research
![PDF] Mass-spectrometry-based spatial proteomics data analysis using pRoloc and pRolocdata | Semantic Scholar PDF] Mass-spectrometry-based spatial proteomics data analysis using pRoloc and pRolocdata | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/e66de323db973521c93ce8f7f60d4494c4934bb8/2-Figure1-1.png)