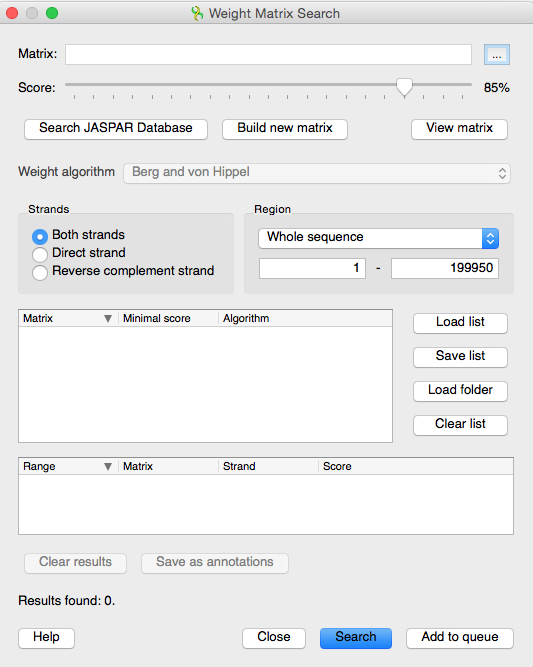
Transcription factor binding sites Prediction. The sequence around the... | Download Scientific Diagram

IMAGE predicts transcription factor binding sites with high confidence.... | Download Scientific Diagram

Prediction of transcription factor bindings sites affected by SNPs located at the osteopontin promoter - ScienceDirect
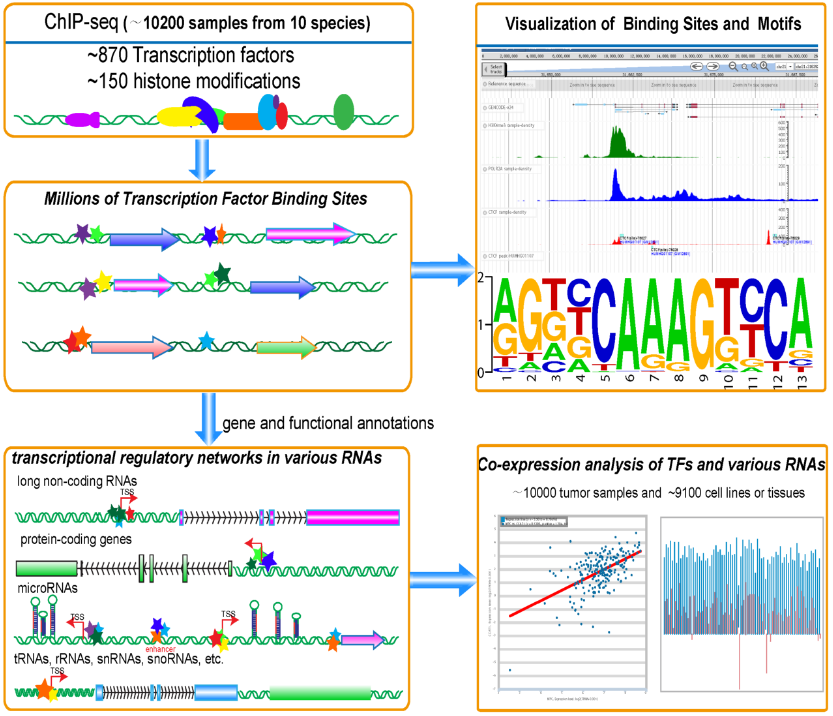
Transcription Factor Binding Sites, Motifs and Expression Profiles from ~10200 ChIP-seq and ~20000 RNA-seq samples
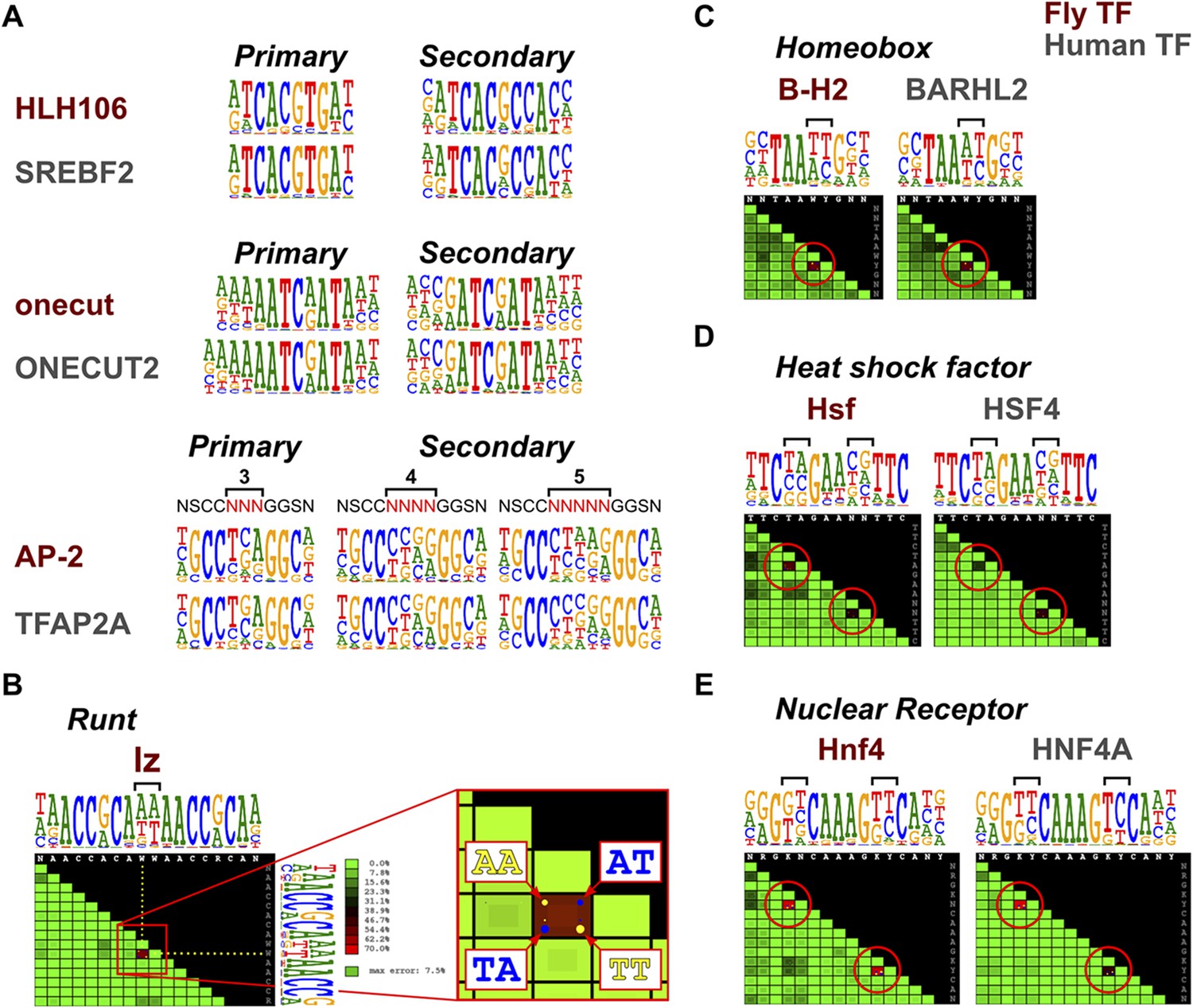
Conservation of transcription factor binding specificities across 600 million years of bilateria evolution | eLife

Prediction of transcription factor binding sites and mutation analysis... | Download Scientific Diagram
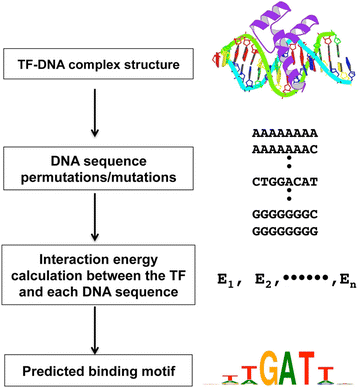
An efficient algorithm for improving structure-based prediction of transcription factor binding sites | BMC Bioinformatics | Full Text

Predicting transcription factor binding sites using DNA shape features based on shared hybrid deep learning architecture: Molecular Therapy - Nucleic Acids
![PDF] DNA Shape Features Improve Transcription Factor Binding Site Predictions In Vivo. | Semantic Scholar PDF] DNA Shape Features Improve Transcription Factor Binding Site Predictions In Vivo. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/5d68bdf0391af25147a48171a54fea49d64d16d4/2-Figure1-1.png)
PDF] DNA Shape Features Improve Transcription Factor Binding Site Predictions In Vivo. | Semantic Scholar
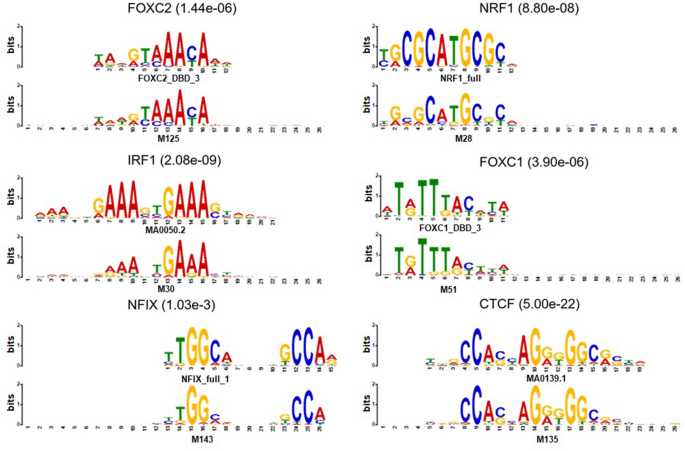
Enhancing the interpretability of transcription factor binding site prediction using attention mechanism | Scientific Reports
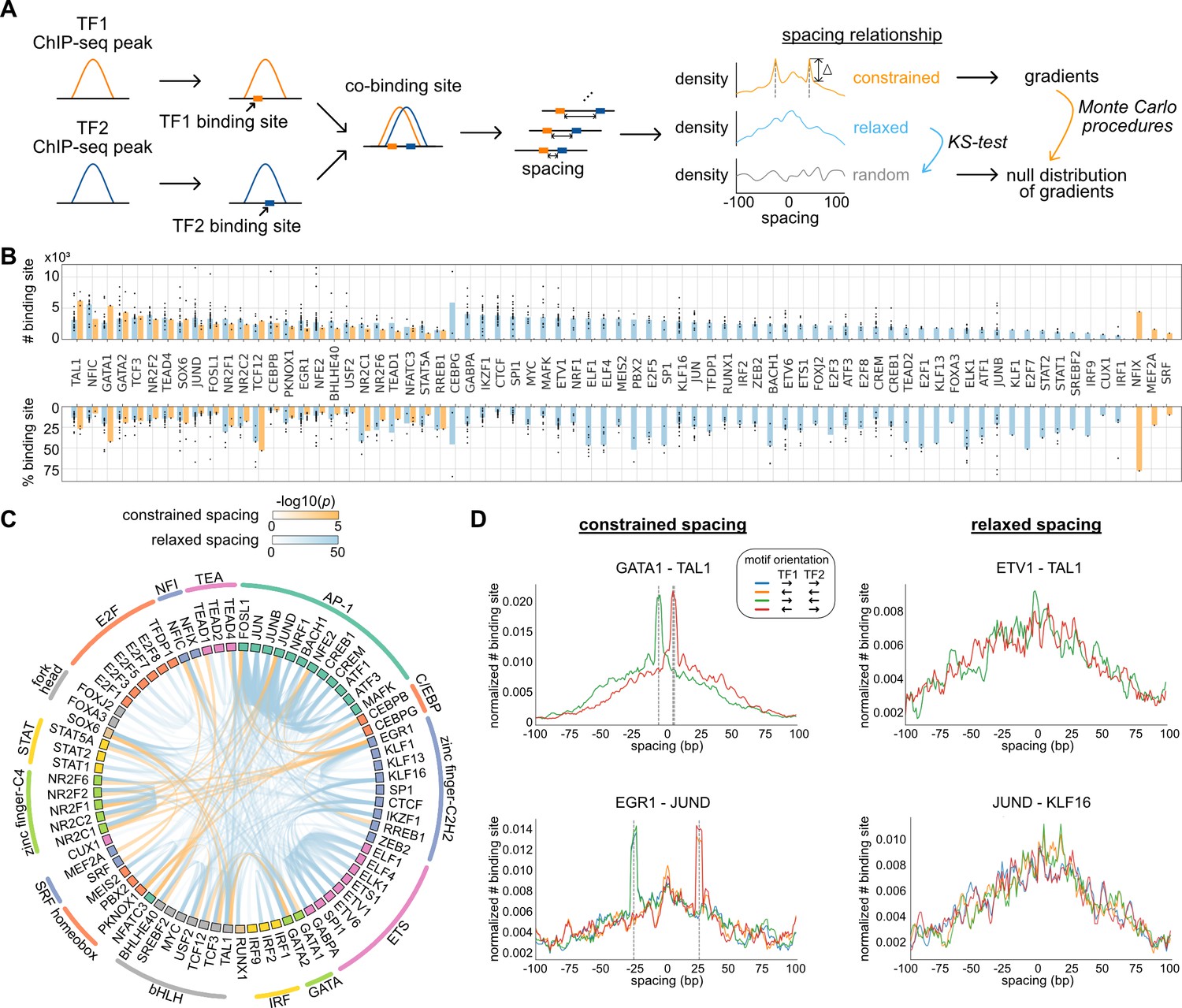
Systematic analysis of naturally occurring insertions and deletions that alter transcription factor spacing identifies tolerant and sensitive transcription factor pairs | eLife